Scientist V
National Institute of Plant Genome Research (NIPGR), New Delhi
- 91-11-26735252, Fax: 91-11-26741658
- vgaur@nipgr.ac.in, vineetgaur1982@gmail.com
Profile
Research Area
Structural Biology, Protein biochemistry, and Biophysics
Research Interest and recent work
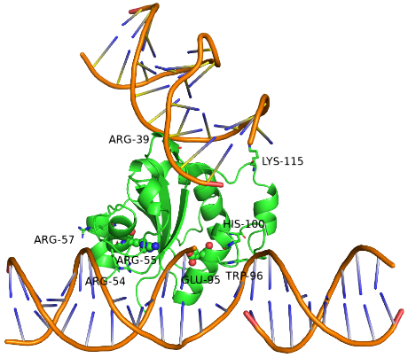
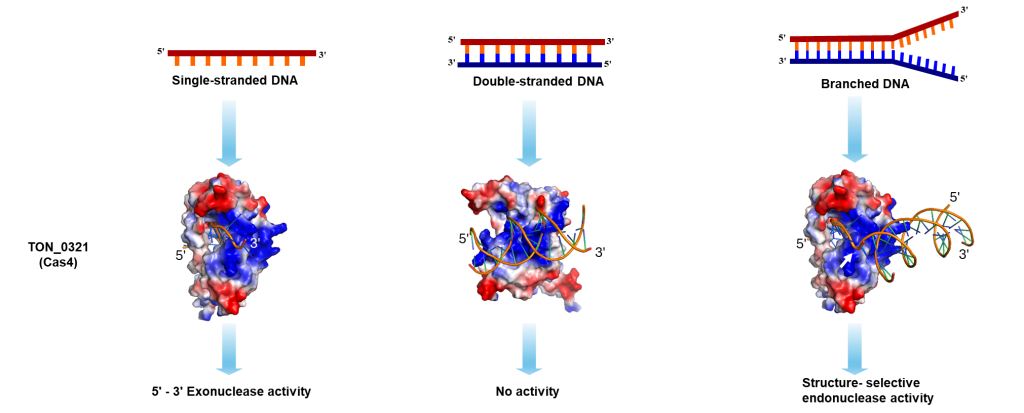
The Laboratory of Structural Biology at NIPGR is interested in understanding various DNA repair and recombination mechanisms in plants from structural and biochemical perspectives. In addition, we are interested in novel nucleases with structure-selective endonuclease activity. We have identified At-HIGLE, a GIY-YIG endonuclease, as a possible plant homolog of Slx1 with Holliday Junction (four-way DNA molecule) resolving potential (left panel figure). We have also identified novel structure-selective endonuclease activity associated with the Cas4 protein from Thermococcus onnurineus (right panel figure).
Professional & Academic Background
Staff Scientist V (2022- present): National Institute of Plant Genome Research, New Delhi.
Ramalingwaswami Fellow (2019-2024): National Institute of Plant Genome Research, New Delhi.
Post-Doctoral Fellow (2012-2019): International Institute of Molecular and Cell Biology, PolandPost-Doctoral Fellow (2012-2019): International Institute of Molecular and Cell Biology, Poland
Post-Doctoral Fellow (2010-2012): The Ohio State University, USA
Ph.D. (2004-2010): National Institute of Immunology, New Delhi
M.Sc. (2002-2004): in Biomedical Sciences from the University of Delhi, New Delhi.
B.Sc. (H) (1999-2002): in Zoology from Hindu College, University of Delhi, New Delhi.
Awards & Honors
Team award of the Polish Minister of Science, Poland (2022).
Core Research Grant, Dept. of Science and Technology, Government of India (2020-2023)
Ramalingaswami Fellowship, Dept. of Biotechnology, Government of India (2019-2024)
NET-JRF (CSIR), Govt. of India (2004-2009).
Graduate Aptitude Test in Engineering (GATE) 2004 with an all-India rank of 20
Catch Them Young scholarship, CSIR-university interaction model (2003-2004)
Jean and Ashit Ganguly Education Scholarship (2002-2004)
Third rank in Delhi University in B.Sc (H) Zoology (2002).
Current Members
Ms. Megha Parihar
Ph.D. student
Mr. Asish Pattnayak
Ph.D. student
Ms. Tamalika Datta
Ph.D. student
Ms. Bharti Sharma
Ph.D. student
Mr. Sreerag Sukumaran
PA-1
Ms. Naiya Chauhan
PA-1
Mr. Ramesh
Lab attendant
Former Members
Ms. Shreya Negi
PA-1
Ms. Reetika Tandon
PA-1
Ms. Poonam Kumari
PA-1
Mr. Praveen Rai
PA-1
Publications
Jain M, Pattnayak AK, Aggarwal S, Rai P, Kavya J, Chandrayan S, Goel M, Gaur V. Branched DNA processing by a thermostable CAS-Cas4 from Thermococcus onnurineus: expanding biochemical landscape of nuclease activity. (2025) J Biol Chem. 110701.
Rai P, Kumari P, Gaur V. Erasing methylation marks on DNA by plant-specific DEMETER family DNA glycosylases. (2025) Journal of Plant Growth Regulation 44 (5), 1810-1826.
Mahtha SK, Kumari K, Gaur V, Yadav G. Cavity architecture based modulation of ligand binding tunnels in plant START domains. (2023) Comput Struct Biotechnol J. 21: 3946-3963.
Kumar U, Goyal P, Madni ZK, Kamble K, Gaur V, Rajala MS, Salunke DM. A structure and knowledge-based combinatorial approach to engineering universal scFv antibodies against influenza M2 protein. (2023) J Biomed Sci. 30: 56.
Jaiswal D, Kumar U, Gaur V, Salunke DM. Epitope-directed anti-SARS-CoV-2 scFv engineered against the key spike protein region could block membrane fusion. (2023) Protein Sci. 32: e4575.
Verma P, Kumari P, Negi S, Yadav G, Gaur V*. Holliday junction resolution by At-HIGLE: an SLX1 lineage endonuclease from Arabidopsis thaliana with a novel in-built regulatory mechanism. (2022) Nucleic Acids Research. 50: 4630–4646
Raul B, Bhattacharjee O, Ghosh A, Upadhyay P, Tembhare K, Singh A, Shaheen T, Ghosh AK, Torres-Jerez I, Krom N, Clevenger J, Udvardi M, Scheffler BE, Akins PO, Sharma RD, Bandyopadhyay K, Gaur V, Kumar S, Sinharoy S. Microscopic and transcriptomic analyses of Dalbergoid legume peanut reveal a divergent evolution leading to Nod Factor dependent epidermal crack-entry and terminal bacteroid differentiation. (2022) Mol Plant Microbe Interact.35: 131-145.
Verma P, Tandon R, Yadav G, Gaur V*. Structural aspects of DNA repair and recombination in crop improvement. (2020) Frontiers in Genetics. 11: 1-30. (* Corresponding author).
Gaur V, Ziajko W, Nirwal S, Szlachcic A, Gapińska M, Nowotny M. Recognition and processing of branched DNA substrates by Slx1-Slx4 nuclease. (2019) Nucleic Acids Research. 47: 11681-11690.
Jaciuk M*, Swuec P*, Gaur V*, Kasprzak JM, Renault L, Dobrychłop M, Nirwal S, Bujnicki JM, Costa A, Nowotny M. A combined structural and biochemical approach reveals translocation and stalling of UvrB on the DNA lesion as a mechanism of damage verification in bacterial nucleotide excision repair. (2019) DNA Repair. 85: 102746. (* Equal contribution).
Nowotny M*, Gaur V*. Structure and mechanism of nucleases regulated by SLX4. (2016) Curr Opin Struct Biol. 36: 97–105. (* Corresponding authors).
Gaur V, Wyatt HD, Komorowska W, Szczepanowski RH, de Sanctis D, Gorecka KM, West SC, Nowotny M. Structural and Mechanistic Analysis of the Slx1-Slx4 Endonuclease. (2015) Cell Reports. 10: 1467–1476.
Gaur V, Vyas R, Fowler JD, Efthimiopoulos G, Feng JY, Suo Z. Structural and kinetic insights into binding and incorporation of L-nucleotide analogs by a Y-family DNA polymerase. (2014) Nucleic Acids Research. 42: 9984–9995.
Espinoza-Herrera SJ, Gaur V, Suo Z, Carey PR. Following DNA chain extension and protein conformational changes in crystals of a Y-family DNA polymerase via Raman crystallography. (2013) Biochemistry 52: 4881-4890.
Tapryal S*, Gaur V*, Kaur KJ, Salunke DM. Structural evaluation of a mimicry-recognizing paratope: plasticity in antigen-antibody interactions manifests in molecular mimicry. (2013) J Immunol. 191:456-63. (* Equal contribution).
Gaur V, Chanana V, Jain A, Salunke DM. The structure of a haemopexin-fold protein from cow pea (Vigna unguiculata) suggests functional diversity of haemopexins in plants. (2011) Acta Cryst. F67: 193-200.
Gaur V, Qureshi IA, Singh A, Chanana V, Salunke DM. Crystal structure and functional insights of hemopexin fold protein from grass pea. (2010) Plant Physiology 152: 1842-1850.
Gupta P, Gaur V and Salunke DM. Purification, identification and preliminary crystallographic studies of a 2S albumin seed protein from Lens culinaris. (2008) Acta Cryst. F64: 733–736.
Gaur V, Sethi DK and Salunke DM. Purification, identification and preliminary crystallographic studies of Pru du amandin, an allergenic protein from Prunus dulcis. (2008) Acta Cryst. F64: 32–35
PDB AND PROTEIN SEQUENCE ENTRIES
3LP9: Crystal structure of LS-24, seed albumin from Lathyrus sativus.
3EHK: Crystal structure of Pru du amandin, an allergenic protein from Prunus dulcis.
3OYO: Crystal structure of hemopexin fold protein CP4 from Vigna unguiculata.
4QW8, 4QW9, 4QWA, 4QWB, 4QWC, 4QWD, 4QWE: Ternary Crystal Structures of a Y-family DNA polymerase Dpo4 from Sulfolobus solfataricus in complex with DNA and dCTP or its L-analogs.
4HOG, 4HOI, 4HOH: Structural evaluation of a mimicry recognizing paratope: plasticity in antigen-antibody interactions manifests in molecular mimicry.
4XLG, 4XM5:Structural and mechanistic analysis of the Slx1-Slx4 endonuclease from Candida glabrata.
6SEH, 6SEI: Recognition and processing of branched DNA substrate by Slx1-Slx4 nuclease from Thermothielavioides terrestris.
7WME: Crystal Structure of the catalytic domain of At-HIGLE from Arabidopsis thaliana.
P86190: Protein sequence of LS-24 from Lathyrus sativus.

