Dr. Naveen Chandra Bisht
Scientist VI, FNA, FASc, FNASc, FNAAS
Ph.D. Dept. of Genetics, Univ. of Delhi South Campus, New Delhi, INDIA
- 91-11-26735183
- ncbisht@nipgr.ac.in, ncbisht@gmail.com
Google Scholar,Research Gate,Vidwan-ID: 53368
ORCID ID: https://orcid.org/0000-0001-7817-2193
Profile
Research Interests
(A) Seed Meal & Oil Quality Enhancement
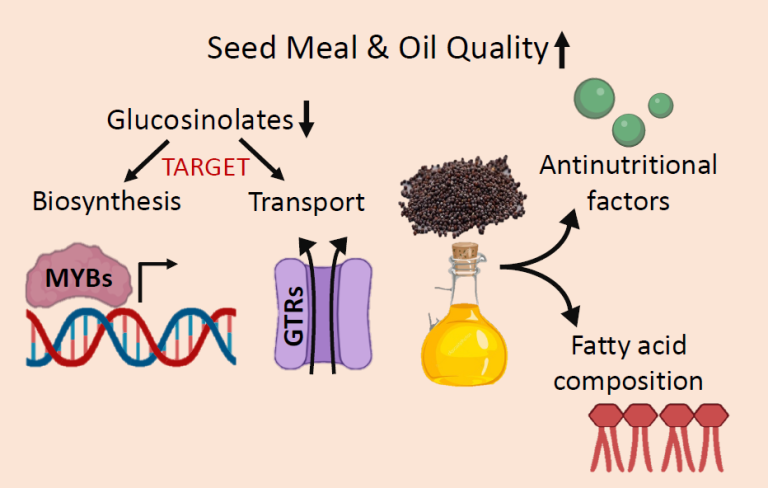
High amount of glucosinolates, erucic acid and several other seed metabolites like phytic acid, sinapine etc. in Brassica oilseeds are considered anti-nutritional and reduces the oil and meal quality. In the quest to improve flavor and nutritional qualities of Brassica oilseed crops, the major thrust area of the laboratory is to understand the genetic and biochemical basis of glucosinolates biosynthesis and transport in Indian oilseed mustard. The lab research focuses on targeting the genes/proteins involved in in the biosynthesis and transport of glucosinolates, fatty acid composition, and other anti-nutritional metabolites through CRISPR/Cas9 and RNAi-based approaches.
(B) Biofortification of Beneficial Glucosinolates
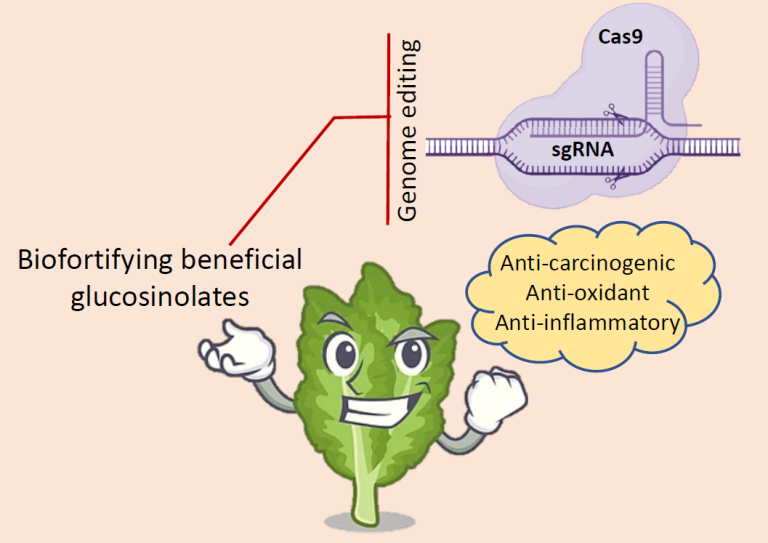
Various glucosinolates and their products also possess anti-carcinogenic, anti-oxidant and anti-inflammatory properties in humans and animals. Metabolic engineering of Indian mustard for enrichment of desirable glucosinolates while reducing the anti-nutritional glucosinolates will improve the food and feed value of this crop. The lab focuses on developing transgene-free Indian mustard varieties with biofortified glucosinolates for human consumption.
(C) Glucosinolates in Plant Defense
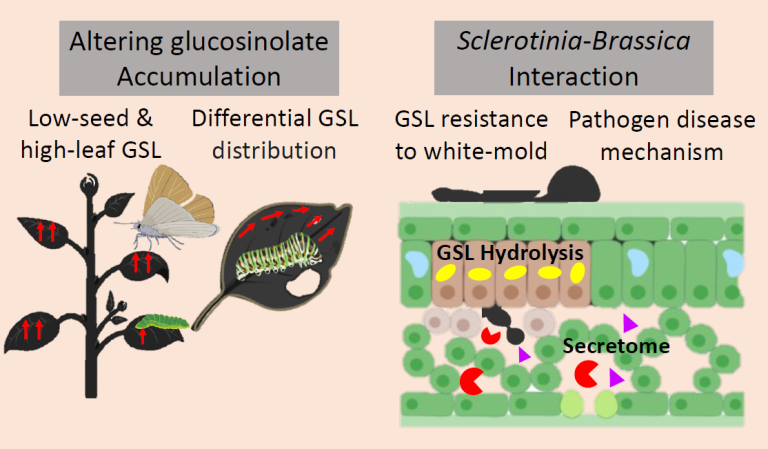
The lab is investigating the role of key players regulating the source-sink relationship in glucosinolate transport process to develop ‘the ideal lines’ with low-seed and high-leaf glucosinolates. Tissue-specific manipulation of glucosinolate accumulation will be a promising strategy to engineer defense against insects and pests of Brassica crops. Additionally, the role of glucosinolate-negotiated defense against the white mold pathogen, Sclerotinia is also being researched. Key pathogen proteins of Sclerotinia-Brassica interaction are being identified and functionally characterized to understand the disease mechanism in-depth
(D) Glucosinolate Catabolism
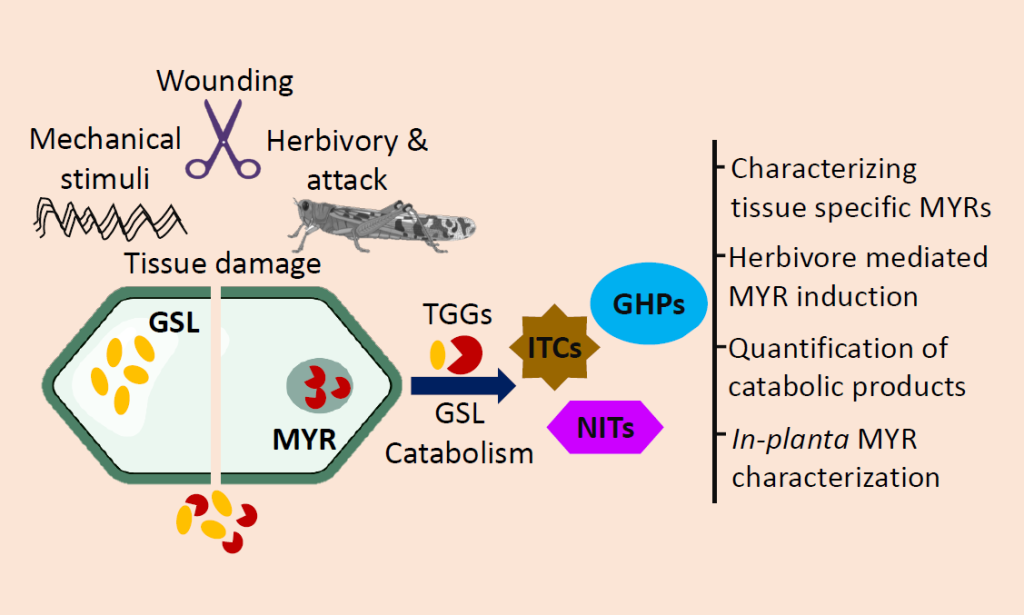
Glucosinolates are a group of amino acid-derived plant secondary metabolites distinctive to the order Capparales, which includes agriculturally important Brassica vegetables, condiments and oilseed crops and the model species Arabidopsis thaliana.
The biological activities of glucosinolates are governed post breakdown by myrosinases (β-thioglucoside glucohydrolase, TGG). Glucosinolate activation by myrosinases upon herbivory is poorly understood in Brassica crops and is further complexed by polyploidy. The lab focuses on in-planta characterization of myrosinase encoding gene family in mustard to understand the regulation of glucosinolate catabolism during herbivory.
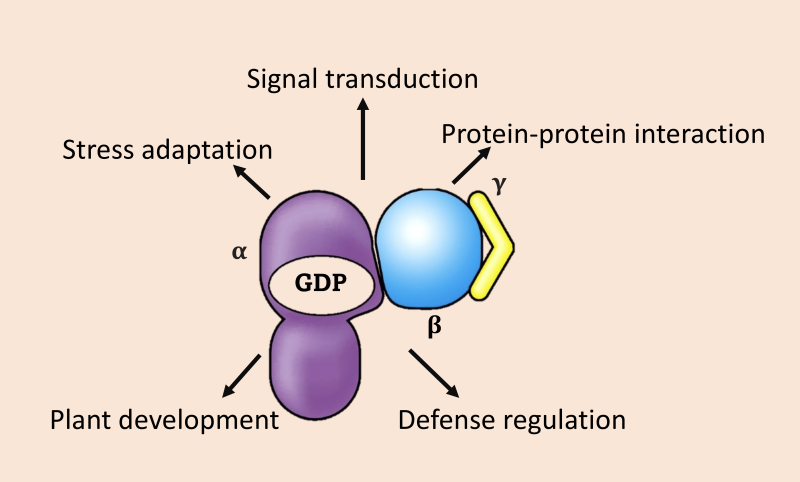
(E) G-proteins in Plant Growth and Defense
The G-protein research in crop species is still in its infancy, and a handful of studies suggest important roles of G-proteins in regulating plant architectural and key agronomical traits including plant’s response to abiotic and biotic factors. The involvement and signal transduction mediated by G-proteins in plant development, stress adaptation and defense regulation are being studies to develop novel approaches to boost plant yield and fitness in the changing environments.
(F) Mechanistic Basis of BCAA Homeostasis in Plants

The branched-chain amino acids (BCAAs) Ile, Val and Leu are essential nutrients that humans and other animals need to obtain from plants. However, the total amounts of plant BCAAs rarely match the human nutritional requirements. Research in our laboratory focuses on understanding the mechanistic basis for BCAA homeostasis and feedback regulation of a key allosteric enzyme (IPMS) of Leu biosynthesis pathway in Brassicas by utilizing an array of evolutionary, biochemical and gene editing tools.
Awards & Honors
Year
Honors
2024
Fellow of the Indian National Science Academy (INSA), New Delhi
2023
Fellow of the Indian Academy of Sciences (IAS), Bangalore, India
2022
Fellow of the National Academy of Agricultural Sciences (NAAS), India
2020
Fellow of the National Academy of Science (NASI), Allahabad, India
2017
Associate of the National Academy of Agriculture Science (NAAS), India
2012
Max Planck India Fellow (2013-2016)
2008
Visiting Scientist at Donald Danforth Plant Science Center, St. Louis, USA through DBT-NIPGR Short Term Overseas Fellowship (June 2008 - July 2010)
Year
Awards
2024
DBT - TATA Innovation Fellowship
2019
DBT - S. Ramachandran National Bioscience Award for Career Developmen
2017
NASI-Scopus Young Scientist Award - Agriculture
2013
DBT-Innovative Young Biotechnologist Award (IYBA)
2007
INSA Young Scientist Medal - Agriculture Biotechnology.
Lab News and Recognition
The Indian Express (August 21, 2023) Gene-edited mustard: Less pungent, more usefulhttps://www.google.com/amp/s/indianexpress.com/article/explained/explained-economics/gene-edited-mustard-less-pungent-more-useful-8901549/lite/
NDTV Profit (August 18, 2023), BQ Prime (August 19, 2023) Delhi Scientists Genome-Edit Mustard Plant To Yield Bland Oil With Insect-Repelling Compoundshttps://www.ndtvprofit.com/nation/delhi-scientists-genome-edit-mustard-plant-to-yield-bland-oil-with-insect-repelling-compounds https://www.google.com/amp/s/www.bqprime.com/amp/nation/delhi-scientists-genome-edit-mustard-plant-to-yield-bland-oil-with-insect-repelling-compounds
ISAAA Gene Editing Supplement (August 16, 2023) CRISPR Mustard Produces Oil with Low Pungency https://www.isaaa.org/kc/cropbiotechupdate/ged/article/default.asp?ID=20357
Lab work highlighted by FEDERATION OF SEED INDUSTRY OF INDIA on Genome Editing for Sustainable Agriculture in India.
YouTube link: https://www.youtube.com/watch?v=FScfpnpLu3E&t=124s
Dr. Roshan Kumar, a former PhD student and a postdoctoral fellow in Dr. Naveen C. Bisht’s lab has been awarded INSA Young Scientist Award 2020 in Agriculture Science. He carried out biochemical and mechanistic studies for the detailed understanding of glucosinolate diversity in Brassica crops.
Lab research on LOW GLUCOSINOLATE MUSTARD highlighted in AGNIi Innovation Showcase Initiative.
YouTube link: https://www.youtube.com/watch?v=Z8N7EWdmHxc&t=12s
Hands of God: Regulations should be in harmony with research for India to progress in gene editing. Cover Story of THE WEEK February 16, 2019 https://www.theweek.in/health/cover/2019/02/16/hands-of-god.html
March 2018: Lab received DBT-BIRAC financial support for event selection trial and compositional analysis of low glucosinolate mustard. The promising low glucosinolate line has been established.
Glucosinolate research of Bisht’s lab received Best Poster Awards in The India International Science Festival for two successive years (2016 & 2017)
Dr. Rehna Augustine work on developing low glucosinolate mustard has received several national accolades including Young Scientist Medal from INSA (2015), NASI (2016) and Jawaharlal Nehru Award (2015) in Plant Biotechnology for outstanding Doctoral Thesis research in Agricultural and Allied Sciences by ICAR, New Delhi.
Co-authored Network
Current Members
Dr. Pravin Kumar
Postdoctoral Fellow : Apr 2018 onwards
Nutritional improvement of Indian oilseed mustard using CRISPR/Cas9-mediated genome editing
- csirpravin@gmail.com
Ms. Bhanu Malhotra
Ph.D. Scholar : Aug 2018 onwards
Investigating the Sclerotinia-Brassica interaction and understanding the glucosinolate-negotiated defense against white-mold
- bhanumalhotra@nipgr.ac.in
Mr. Mohan Varghese
Ph.D. Scholar : Aug 2018 onwards
Investigating the mechanistic basis of Leucine biosynthesis in plants
- mohan_tnau@nipgr.ac.in
Ms. Avni Mann
Ph.D. Scholar : Aug 2019 onwards
Understanding the genetic basis of tissue-specific glucosinolate diversity in Indian oilseed mustard
- avnimann@nipgr.ac.in
Ms. Kajal
Ph.D. Scholar : Sept 2021 onwards
Improvement of fatty acid composition in Indian oilseed mustard
- kajalmittal@nipgr.ac.in
Ms. Anjala K.
Ph.D. Scholar : Aug 2022 onwards
Nutritional improvement of phytic acid trait in oilseed Brassica
- anjala@nipgr.ac.in
Mr. Shailendra Kharwal
Technical Officer : july 2017 onwards
Lab responsibilities: Overall management of various lab resources
- shail2385@nipgr.ac.in
Mr. Vinod Kumar
Multitasking Staff
Lab responsibilities: Trained workforce for glucosinolate analysis of Brassica germplasm and experimental field management
- vinodkumar868@gmail.com
Mr. Amal Roul
Project Technical Assistant
Lab responsibilities: Skilled workforce for various activities including autoclaving, soil preparation and lab hygiene
- amalroul555@gmail.com
Mr. Hari Bhagvan Maurya
NIPGR Field Assistant
Lab responsibilities: Lab technician experienced in sample harvesting, recording and managing data, and experimental field management
- haribhagvanmaurya5548@gmail.com
Mr. Raju Das
NIPGR Field Assistant
Lab responsibilities: Maintenance of Brassica germplasm in green house and experimental fields
Former Group Members
Dr. Pawan Kumar
PhD and Postdoctoral Fellow
(2015 – 2023)
Dr. Sonam Chaudhary
DBT-RA & SERB-NPDF
(2019 – 2025)
Ms. Ruchi Tiwari
Ph.D. & Postdoctoral Fellow
(2016 – 2024)
Ms. Praveena Kanchupati-Challa
Ph.D.-JRF
(2009-2011)
Dr. Meenu
Postdoctoral Fellow
(2009-2011)
Dr. Ruby Chandna
Postdoctoral Fellow
(2011-2012)
Ms. Ishita
Project Fellow (JRF)
(2013-2014)
Dr. Rehna Augustine
Ph.D. & Postdoctoral Fellow
(2007-2017)
Dr. Gulab C. Arya
Ph.D. & Postdoctoral Fellow
(2011-2018)
Dr. Prabodh Bajpai
Postdoctoral Fellow (NPDF)
(2016-2018)
Dr. Deepti M. Nambiar
Ph.D. Scholar
(2012-2019)
Dr. Roshan Kumar
Ph.D. & Postdoctoral Fellow
(2011-2020)
Ms. Oshin Sharma
Project Fellow (JRF)
(2019-2020)
Dr. Bornali Gohain
Postdoctoral Fellow
(2019-2021)
Ms. Aprajita Sharma
Project Fellow (SRF)
(2018-2021)
Dr. Juhi Kumari
Ph.D. Scholar
(2015-2022)
Complete List of Publications
Patents
Kumari J, Mann A, Kumar R, Bisht NC. Recombinant expression cassettes for modification of glucosinolate content in plants. Indian Patent application no. 202211034971 (filed 17.06.22); PCT no. WO 2023/242872 (Published on 21.12.2023); Australian Patent application no. 2023290708 (filed 29.11.2024); Canadian Patent application no. 3259474 (filed 13.12.2024).
Bisht NC and Augustine R. Compositions and methods for production of transgenic plants having reduced glucosinolate levels. Indian Patent 411478 (granted on 15.11.2022)
Bisht NC, Jagannath A, Gupta V, Burma PK and Pental D. A novel method for obtaining improved fertility restorer lines for transgenic male-sterile crop plants and a DNA construct for use in said method. US Patent 7741541 (granted on 22.06.2010).
Bisht NC, Jagannath A, Gupta V, Burma PK and Pental D. A novel method for obtaining improved fertility restorer lines for transgenic male-sterile crop plants and a DNA construct for use in said method. European Patent 1644506 (granted on 09.09.2009).
Bisht NC, Jagannath A, Gupta V, Burma PK and Pental D. A novel method for obtaining improved fertility restorer lines for transgenic male-sterile crop plants and a DNA construct for use in said method. Indian Patent 238973 (granted on 03.03.2010).
Publications (in chronological order)
Kumar P, Bisht NC (2025) High-level production of health-beneficial glucoraphanin by multiplex editing of the AOP2 gene family in mustard. Plant Biotechnology Journal (doi/10.1111/pbi.70171)
Varghese M, Kumar R, Sharma A, Lone A, Gershenzon J, Bisht NC (2025) Isopropylmalate synthase regulatory domain removal abolishes feedback regulation at the expense of leucine homeostasis in plants. Plant Physiology 197: kiaf041
Maji S, Waseem M, Sharma MK, Singh M, Singh A, Dwivedi N, Thakur P, Cooper DG, Bisht NC, Fassler JS, Subbarao N, Khurana JP, Bhavesh NS, Thakur JK (2024) MediatorWeb: a protein-protein interaction network database for the RNA polymerase II Mediator complex. FEBS Journal 291: 3938-3960
Tiwari R, Garg K, Senthil-Kumar M, Bisht NC (2024) XLG2 and CORI3 function additively to regulate plant defense against the necrotrophic pathogen Sclerotinia sclerotiorum. The Plant Journal 117: 616-631
Naik J, Tyagi S, Rajput R, Kumar P, Pucker B, Bisht NC, Misra P, Stracke R, Pandey A (2024) Flavonols have opposite effects on the interrelated glucosinolate and camalexin biosynthetic pathways in Arabidopsis thaliana. Journal of Experimental Botany 75: 219-240
Avni M, Juhi K, Roshan K, Pawan K, Pradhan AK, Pental D, Bisht, NC (2023) Targeted editing of multiple homologs of GTR1 and GTR2 genes provides the ideal low-seed, high-leaf glucosinolate oilseed mustard with uncompromised defense and yield. Plant Biotechnology Journal 21: 2182-2195 (Cover image of PBJ Volume 21, Issue 11, Nov 2023)
Malhotra B, Kumar P, Bisht NC (2023) Defense versus growth trade-offs: Insights from glucosinolates and their catabolites. Plant Cell & Environment 46: 2964-2984.
Sharma A, Sinharoy S, Bisht NC (2023) The mysterious non-arbuscular mycorrhizal status of Brassicaceae species. Environmental Microbiology 25: 917-930
Kumar R, Reichelt M, Bisht NC (2022) An LC-MS/MS assay for enzymatic characterization of methylthioalkylmalate synthase (MAMS) involved in glucosinolate biosynthesis. Methods in Enzymology 676: 49-69
Dwivedi V, Parida SK, Chattopadhyay D (2017) A repeat length variation in myo-inositol monophosphatase gene contributes to seed size trait in chickpea. Scientific Reports. 7:4764
Kumar R, Bisht NC (2022) Interacting partners of Brassica juncea Regulator of G-protein Signaling protein suggest its role in cell wall metabolism and cellular signaling. Bioscience Reports 42: BSR20220302
Tiwari R, Bisht NC (2022) The multifaceted roles of heterotrimeric G-proteins: lessons from models and crops. Planta 255, 88
Tiwari R, Kaur J, Bisht NC (2021) Extra-large G-proteins influence plant response to Sclerotinia sclerotiorum by regulating glucosinolate metabolism in Brassica juncea. Molecular Plant Pathology 22: 1180-1194
Arya GC, Tiwari R, Bisht NC (2021) A complex interplay of Gβ and Gγ proteins regulates plant growth and defence traits in the allotetraploid Brassica juncea. Plant Molecular Biology 106, 505-520
Nambiar DM, Kumari J, Augustine R, Kumar P, Bajpai PK, Bisht NC (2021) GTR1 and GTR2 transporters differentially regulate tissue-specific glucosinolate contents and defence responses in the oilseed crop Brassica juncea. Plant, Cell & Environment 44, 2729-2743
Gohain B, Kumar P, Malhotra B, Augustine R, Pradhan AK, Bisht NC (2021) A comprehensive Vis-NIRS equation for rapid quantification of seed glucosinolate content and composition across diverse Brassica oilseed chemotypes. Food Chemistry 354, 129527
Kumar P, Yadava S, Singh P, Bhayana L, Mukhopadhyay A, Gupta V, Bisht NC, Zhang J, Kudrna D, Copetti D, Wing RA, Reddy VB, Pradhan AK, Pental D (2021) A chromosome-scale assembly of allotetraploid Brassica juncea (AABB) elucidates comparative architecture of the A and B genomes. Plant Biotechnology Journal 19, 602-614
Malhotra B, Bisht NC (2020) Editorial: Glucosinolates: Regulation of Biosynthesis and Hydrolysis. Frontiers in Plant Science 11, 620965
Kumar R, Bisht NC (2020) Heterotrimeric Gα subunit regulates plant architecture, organ size and seed weight in the oilseed Brassica juncea. Plant Molecular Biology 104, 549-560
Nambiar DM, Kumari J, Arya GC, Singh AK, Bisht NC (2020) A cell suspension based uptake method to study high affinity glucosinolate transporters. Plant Methods 16, 75
Kumar R, Lee SL, Augustine R, Reichelt M, Vassão DG, Palavalli MH, Allen A, Gershenzon J, Jez JM, Bisht NC (2019) Molecular basis of the evolution of methylthioalkylmalate synthase and diversity of methionine-derived glucosinolates. The Plant Cell 31: 1633-47
Bajpai PK, Reichelt M, Augustine R, Gershenzon J, Bisht NC (2019). Heterotic patterns of primary and secondary metabolites in the oilseed crop Brassica juncea. Heredity 123: 318-36
Augustine R, Bisht NC (2019) Targeted silencing of genes in polyploids: lessons learned from Brassica juncea-glucosinolate system. Plant Cell Reports 38: 51-57
Kumar R, Bisht NC (2018) Duplicated RGS (Regulator of G-protein signaling) proteins exhibit conserved biochemical but differential transcriptional regulation of heterotrimeric G-protein signaling in Brassica species. Scientific Reports 8: 2176
Kumar P, Augustine R, Singh AK, Bisht NC (2017) Feeding behaviour of generalist pests on Brassica juncea: implication for manipulation of glucosinolate biosynthesis pathway for enhanced resistance. Plant, Cell & Environment 40: 2109-20
Kumar R, Sharma A, Chandel I, Bisht NC (2017) Pattern of expression and interaction specificity of multiple G-protein beta (Gβ) subunit isoforms with their potential target proteins reveal functional dominance of BjuGβ1 in the allotetraploid Brassica juncea. Plant Physiology and Biochemistry 118: 22-30
Pandey C, Augustine R, Panthri M, Zia I, Bisht NC, Gupta M (2017) Arsenic affects the production of glucosinolate, thiol and phytochemical compounds: A comparison of two Brassica cultivars. Plant Physiology and Biochemistry 111: 144-154
Chandna R, Augustine R, Kanchupati P, Kumar R, Kumar P, Arya GC, Bisht NC (2016) Class-specific evolution and transcriptional differentiation of 14-3-3 family members in mesohexaploid Brassica rapa. Frontiers in Plant Science 7: 12
Gupta SA, Arya GC, Malviya N, Bisht NC and Yadav DK (2016) Molecular cloning and expression profiling of multiple Dof genes of Sorghum bicolor (L) Moench. Molecular Biology Reports 43: p767-74
Augustine R, Bisht NC (2015) Biofortification of oilseed Brassica juncea with the anti-cancer compound glucoraphanin by suppressing GSL-ALK gene family. Scientific Reports 5: 18005
Augustine R, Bisht NC (2015) Biotic elicitors and mechanical damage modulate glucosinolate accumulation by co-ordinated interplay of glucosinolate biosynthesis regulators in polyploid Brassica juncea. Phytochemistry 117: 43-50
Meenu, Augustine R, Majee M, Pradhan AK, Bisht NC (2015) Genomic origin, expression differentiation and regulation of multiple genes encoding CYP83A1, a key enzyme for core glucosinolate biosynthesis, from the allotetraploid Brassica juncea. Planta 241: 651-665
Bisht NC, Jagannath A, Augustine R, Burma PK, Gupta V, Pradhan AK, Pental D (2015) Effective restoration of male-sterile (barnase) lines requires overlapping and higher levels of barstar expression: A multi-generation field analysis in Brassica juncea. Journal of Plant Biochemistry and Biotechnology 24: 393-399
Malviya N, Gupta S, Singh VK, Yadav MK, Bisht NC, Sarangi BK, Yadav D (2015) Genome wide in silico characterization of Dof gene families of pigeonpea (Cajanus cajan (L) Millsp.). Molecular Biology Reports 42: 535-552
Gupta S, Malviya N, Kushwaha H, Nasim J, Bisht NC, Singh VK, Yadav D (2015) Insights into structural and functional diversity of Dof (DNA binding with one finger) transcription factor. Planta 241: 549-562
Kumar R, Arya GC, Bisht NC (2014) Differential expression and interaction specificity of the heterotrimeric G-protein family in Brassica nigra reveal their developmental- and condition-specific roles. Plant and Cell Physiology 55:1954-68
Malhotra B, Bisht NC (2020) Editorial: Glucosinolates: Regulation of Biosynthesis and Hydrolysis. Frontiers in Plant Science 11, 620965
Arya GC, Kumar R, Bisht NC (2014) Evolution, expression differentiation and interaction specificity of heterotrimeric G-protein subunit gene family in the mesohexaploid Brassica rapa. PLoS One. 9: e105771
Augustine R, Arya GC, Nambiar DM, Kumar R, Bisht NC (2014) Translational genomics in Brassica crops: challenges, progress, and future prospects. Plant Biotechnology Reports 8: 65-81
Gupta S, Kushwaha H, Singh VK, Bisht NC, Sarangi BK and Yadav D (2014) Genome wide in silico characterization of Dof transcription factor gene family of sugarcane and its comparative phylogenetic analysis with Arabidopsis, rice and sorghum. Sugar Technology 16: 372-384
Augustine R, Majee M, Gershenzon J, Bisht NC (2013) Four genes encoding MYB28, a major transcriptional regulator of aliphatic glucosinolate pathway are differentially expressed in the allopolyploid Brassica juncea. Journal of Experimental Botany 64: 4907-21
Augustine R, Mukhopadhyay A, Bisht NC (2013) Targeted silencing of BjMYB28 transcription factor gene directs development of low glucosinolate lines in oilseed Brassica juncea. Plant Biotechnology Journal 11: 855-66
Kushwaha H, Gupta S, Singh VK, Bisht NC, Sarangi BK, Yadav D (2013) Cloning, in silico characterization and prediction of three-dimensional structure of SbDof1, SbDof19, SbDof23 and SbDof24 proteins from sorghum [Sorghum bicolor (L.) Moench]. Molecular Biotechnology 54: 1-12
Chandna R, Augustine R, Bisht NC (2012) Evaluation of candidate reference genes for gene expression normalization in Brassica juncea using real time quantitative RT-PCR. PLoS One e36918
Choudhury SR, Westfall CS, Laborde JP, Bisht NC, Jez JM, Pandey S (2012) Two chimeric Regulator of G-protein Signaling (RGS) proteins differentially regulate soybean heterotrimeric G-protein cycle. Journal of Biological Chemistry 287: 17870-81
Korekar G, Sharma RK, Kumar R, Meenu, Bisht NC, Srivastava RB, Ahuja PS, Stobdan T (2012) Identification and validation of sex-linked SCAR markers in dioecious Hippophae rhamnoides L. (Elaeagnaceae). Biotechnology Letters 34: 973-8
Bisht NC, Jez JM, Pandey S (2011) An elaborate heterotrimeric G-protein family from soybean expands the diversity of plant G-protein networks. New Phytologist 190: 35-48
Choudhury SR, Bisht NC, Thompson R, Todorov O, Pandey S (2011) Conventional and novel Gγ protein families constitute the heterotrimeric G-protein signaling network in soybean. PLoS One 6: e23361
Guttikonda SK, Trupti J, Bisht NC, Chen H, An C, Pandey S, Xu D, Yu O (2010) Whole genome co-expression analysis of soybean cytochrome P450 genes identifies nodulation-specific P450 monooxygenases. BMC Plant Biology 10: 243
Bisht NC, Gupta V, Ramchiary N, Sodhi YS, Mukopadhyay A, Arumugam N, Pental D, Pradhan AK (2009) Fine mapping of loci involved with glucosinolate biosynthesis in oilseed mustard (Brassica juncea) using genomic information from allied species. Theoretical and Applied Genetics 118: 413-21
Panjabi P, Jagannath A, Bisht NC, Padmaja L, Sharma S, Gupta V, Pradhan AK, Pental D (2008) Comparative mapping of Brassica juncea and Arabidopsis thaliana using Intron Polymorphism (IP) markers: homeologous relationships, diversification and evolution of the A, B and C Brassica genomes. BMC Genomics 9: 113
Ramchiary N#, Bisht NC#, Gupta V#, Mukopadhyay A#, Arumugam N#, Sodhi YS, Pental D, Pradhan AK* (2007) QTL analysis reveals context-dependent loci for seed glucosinolate trait in oilseed Brassica juncea: Importance of recurrent selection backcross scheme for the identification of ‘true’ QTL. Theoretical and Applied Genetics 116: 77-85 (equal contribution)
Bisht NC, Jagannath A, Burma PK, Pradhan AK, Pental D (2007) Retransformation of a male sterile barnase line with the barstar gene as an efficient alternative method to identify male sterile-restorer combinations for heterosis breeding. Plant Cell Reports 26: 727-33
Ray K, Bisht NC, Pental D, Burma PK (2007) Development of barnase/barstar transgenics for hybrid seed production in Indian oilseed mustard (Brassica juncea L. Czern & Coss) using a mutant acetolactate synthase gene conferring resistance to imidazolinone-based herbicide ‘Pursuit’. Current Science 93: 1390-96
Bisht NC, Jagannath A, Gupta V, Burma PK, Pental D (2004) A two gene-two promoter system for enhanced expression of a restorer gene (barstar) and development of improved fertility restorer lines for hybrid seed production in crop plants. Molecular Breeding 14:129-44
Bisht NC, Burma PK, Pental D (2004) Development of 2,4-D resistant lines in Indian mustard (Brassica juncea). Current Science 87: 367-70
Books authored
Malhotra B, Tiwari R, Bisht NC (2024). A guide to culturing, maintenance and leaf inoculation methods for rapid screening and quantification of Sclerotinia sclerotiorum infection in mustard. In M. Senthil-Kumar & H. Ramanna (Eds.), Plant Biotic Stress (pp. XX-XX). Methods in Molecular Biology. Springer Humana Press. New York. (in press)
Varghese M, Malhotra B, Bisht NC (2022) Genome Editing in Polyploid Brassica Crops. In: Kole C, Mohapatra T (eds) The Brassica juncea Genome. Compendium of Plant Genomes. Springer, Cham. pp 471-491 https://doi.org/10.1007/978-3-030-91507-0_25
Bisht NC and Augustine R (2019) Development of Brassica Oilseed Crops with Low Antinutritional Glucosinolates and Rich in Anticancer Glucosinolates. In: Pawan Kumar Jaiwal et al. (eds) Nutritional Quality Improvement in Plants: Concepts and Strategies in Plant Sciences. Springer Press, Switzerland. pp271-287
Augustine R and Bisht NC (2017) Regulation of Glucosinolate Metabolism: From Model Plant Arabidopsis thaliana to Brassica Crops. In: Ramawat K, Mérillon J-M (eds) Reference Series in Phytochemistry: Glucosinolates. Springer Press, Germany. pp163-199
Yadav D*, Anand G, Yadav S, Dubey AK, Bisht NC and Sarangi BK (2013) Intellectual Property Rights in Plant Biotechnology: Relevance, Present Status and Future Prospects. In: Barh D (eds.) OMICS Applications in Crop Sciences. CRC Press, Taylor & Francis Group. pp621-670
Gupta V, Pradhan AK, Bisht NC, Sodhi YS, Arumugam N, Mukopadhyay A and Pental D*. Mapping and tagging of agronomically important genes in Brassica juncea. In: Proceedings 12th International Rapeseed Congress: Sustainable Development in Cruciferous Oilseed Crops Production, March 26-30, 2007; Wuhan China, Vol 1: pp298-300.

