Scientist III
Postdoctoral fellow: Umeå, Plant Science, Center, Umeå, Sweden
PhD: Leibniz Institute of Plant Biochemistry, Halle (Saale), Germany
- 91-11-26741612,14,17 Ext. - 104
- shiv@nipgr.ac.in
Profile
Research Interests
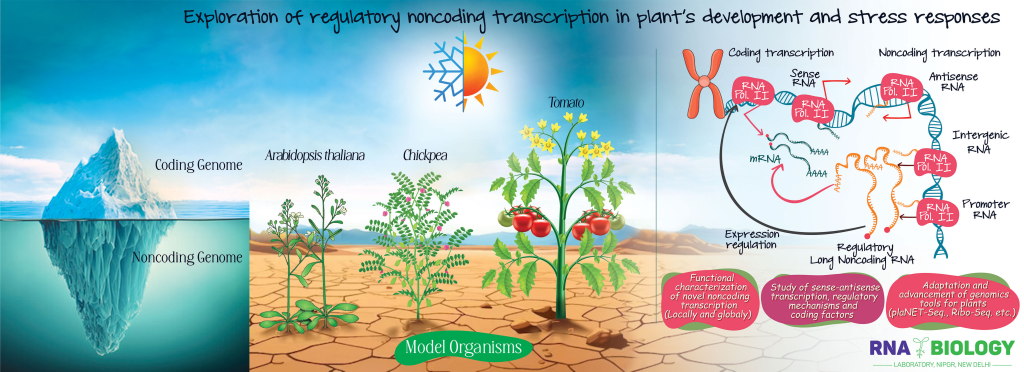
Functional characterization of the widespread noncoding transcription in plants
One of the most important sets of discoveries in the past two decades is the illumination of the ‘dark matter’ of the genome. The advent of novel sequencing technologies could result in showing that most eukaryotic genomes undergo pervasive transcription, comprising more than ~95% of the transcriptome as noncoding RNA. These noncoding RNA transcripts can be of various types such as housekeeping small RNAs, t-RNA, r-RNA, snoRNA, small regulatory RNAs, and long noncoding RNAs (lncRNAs), among others. A substantial portion of this noncoding transcriptome is made up of lncRNAs, the non-protein coding RNA molecules that are longer than 200 nucleotides in length and can be transcribed by RNA Pol II from genomic regions categorized as promoters, introns, intergenic DNA, or the antisense DNA strand. lncRNAs have emerged as key regulators in various fundamental biological processes, including growth, development, disease, and stress responses in both animals and plants. lncRNAs can regulate expression of target gene directly or indirectly through a variety of mechanisms, e.g., via target mimicry of miRNAs, R-loop formation, transcriptional collisions, RNA silencing, etc. A growing number of evidence(s) indicates that the functional and mechanistic exploration of lncRNAs will not only unlock the mysteries of the ‘dark matter’ of the genome but also pave the way for novel findings and noncoding RNA-based applications in agriculture and biotechnology.
Although high-throughput sequencing studies have enabled the identification of tens of thousands of lncRNAs, only a tiny fraction of lncRNAs have been experimentally characterized in plants. Notably, in comparison to the relative progress made in animal systems, our understanding of the roles of lncRNAs, or noncoding transcription in general, their molecular modes of actions, and general RNA mediated regulatory mechanisms involving lncRNAs (if any), in plants is just beginning.
Some of the primary research directions in our RNA biology group at NIPGR, New Delhi include investigations into i) the identification and functional characterization of novel regulatory lncRNAs, ii) study of biologically important noncoding RNA mediated genome-wide transcriptional behaviors, and iii) investigations into whether lncRNAs adapt general regulatory mechanisms, for example, antisense lncRNAs as a class iv) application, adaptation and advancement of novel genomics tools in plants.
We employ diverse molecular genetic, RNA biology, biochemical, biophysical, and high-throughput sequencing approaches in our experiments. Our studies emphasize on development, abiotic and biotic stress conditions. We particularly focus on using transient cold (frost), heat, and drought as model stresses in Arabidopsis thaliana, chickpea, and tomato. In the long run, we ultimately aim to generate molecular understandings of noncoding transcription and lncRNA roles in plants that can be exploited in applied agriculture and in the development of climate-resilient crops.
Study of sense-antisense transcription, antisense lncRNAs, and overlapping protein-coding factors
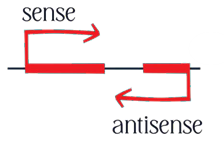
- Characterization of the roles of antisense lncRNAs that overlap with protein-coding genes, and the associated molecular mechanism(s) of action.
- Investigation into the local and global transcriptional dynamics of widespread sense-antisense transcription, and the regulatory processes that govern optimal non-coding and coding transcriptional output during developmental and stress responses in plants.
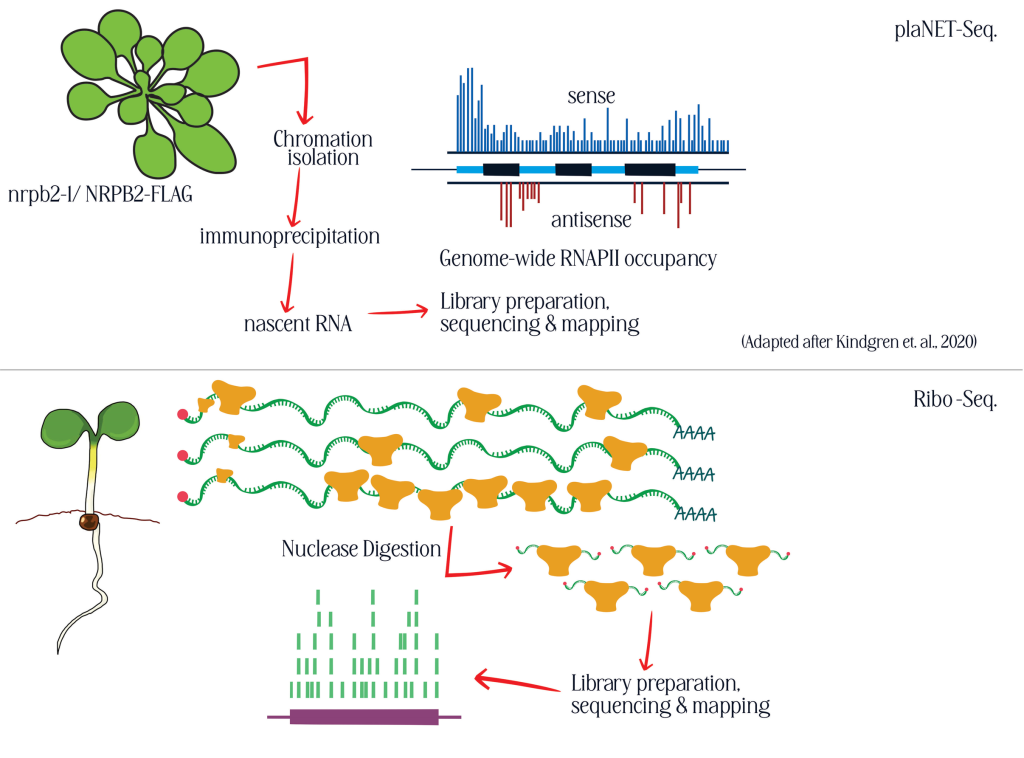

Due to the climate change, the future weather seasons are suspected to be extreme and unpredictable including high frequency severe spring and winter frosts, unpredicted drought and other abiotic stresses and thereby posing a major challenge to crops and sustainable agriculture. lncRNAs are shown to be crucial for both development and stress responses and potentially hold the key for plant’s ability to withstand against harsh weather conditions.By understanding and harnessing the regulatory roles of lncRNAs, we aim to develop crop varieties with improved tolerance to abiotic stress conditions such as frost and drought.
Professional & Academic Background
Staff Scientist III (2023- Present) : National Institute of Plant Genome Research, New Delhi
Post-Doctoral Fellow (2021-2023) : Umeå Plant Science, Center, Umeå, Sweden
Post-Doctoral Fellow (2020-2021) : Leibniz Institute of Plant Biochemistry, Halle (Saale), Germany
Ph.D (2014-2020) : Leibniz Institute of Plant Biochemistry, Halle (Saale),Germany/Martin-Luther-University Halle-Wittenberg, Germany
Awards & Honors
Prime Minister's Early Career Research Grant, 2025, ANRF, India
Carl Tryggers Foundation Postdoctoral Fellowship (2021), UPSC, Umeå, Sweden
Graduiertenkolleg (Deutsche Forschungsgemeinschaft [GRK-1591], German Research Foundation) 2019
Erasmus Mundus Doctoral Scholarship (2014) European Union (Germany)
National Institute of Immunology, New Delhi, India, Fellowship (2010)
Qualified Graduate Aptitude Test in Engineering [GATE] (2010)
Junior Research Fellowship [JRF-NET, 2009], Council of Scientific and Industrial Research, India
Current Members
Dr. Jagajjit Sahu
Postdoctoral Scientist
Tota Mondal
Ph.D student
Gourav Maharia
Ph.D student
Neha
Ph.D student
Manobika Das
Visiting student
Nitya Tiwari
Project Associate
Akshita Nayak
Project Associate
Rashmi Rani Boro
Project Associate
Alumni
Ms. Sanya Bajaj
Dissertation Trainee
Guru Jambheshwar University of Science and Technology, Hissar, Haryana, India
Mr. Raman Aind
Summer Trainee
Delhi Technological University, Delhi, India
Mr. Jagadeep Brar
Summer Trainee
Hansraj College, Delhi University, India
Ms. Harshita Ojha
Dissertation Trainee
Mohanlal Sukhadia University, Udaipur, Rajasthan, India
Ms. Urvija Agrawal
Summer Trainee
Indian Institute of Science, Education and Research, Pune, India
Ms. Prachi Joshi
Dissertation Trainee
Dr. Bhimrao Ambedkar University, Agra, Uttar Pradesh, India
Publications
Zacharaki V., Quevedo M., Nardeli S.M., Meena S.K., Monte E., and Kindgren P. (2025). Convergent antisense transcription primes hosting genes for stress responsiveness in plants. Molecular Plant, doi: https://doi.org/10.1016/j.molp.2025.10.001
Shiv Kumar Meena, Marti Quevedo, Sarah Muniz Nardeli, Clément Verez, Susheel Sagar Bhat, Vasiliki Zacharaki, Peter Kindgren, Antisense transcription from stress-responsive transcription factors fine-tunes the cold response in Arabidopsis, The Plant Cell, 2024;, koae160
Vasiliki Zacharaki, Shiv Kumar Meena, Peter Kindgren, The non-coding RNA SVALKA locus produces a cis- natural antisense transcript that negatively regulates the expression of CBF1 and biomass production at normal temperatures, Plant Communications, 2023, 100551
Meena, S.K., Heidecker, M., Engelmann, S., Jaber, A., de Vries, T., Triller, S., Baumann-Kaschig, K., Abel, S., Behrens, S.-E. and Gago-Zachert, S. (2022), Altered expression levels of long noncoding natural antisense transcripts overlapping the UGT73C6 gene affect rosette size in Arabidopsis thaliana. Plant J
Meena, S. K., Wagner, C., Caggegi, L., Baumann-Kaschig, K. and Ried, M. K. (2021). A user- friendly protocol for the cultivation and successful crossing of Lotus japonicus. Bio-protocol
Srivastava A*, Meena SK*, Alam M, Nayeem SM, Deep S, Sau AK (2013) Structural and functional insights into the regulation of Helicobacter pylori arginase activity by an evolutionary non-conserved motif. Biochemistry 2013 52(3), 508-519. (*-Co first author)
Are you insterested to become a member of our plant RNA Biology lab at NIPGR
We are continuously looking for ‘genuinely motivated’ colleagues with diverse backgrounds who wish to gain experience(s) and/or willing to contribute in any of the above research directions. You can join us as postdoctoral fellow, PhD student or M.Sc./MTech/B.Sc./B. Tech thesis/dissertation trainee(s).
Postdoctoral fellows:
Experienced researchers can contact the group leader directly with very well-structured plans synchronized with a putative time frame along with potential funding options. Prospective postdoc applicants may choose any of the above research theme(s) or can propose their own innovative idea(s) with overlapping/complementary research interests. Interested postdoc candidate(s) may please contact PI by sending a detailed motivation letter and CV explaining their suitability to the group, previous expertise, vision and future goals along with details of 1-3 references.
Ph.D. students/project associates:
PhD admissions takes place via NIPGR, New Delhi PhD program each year which requires the candidates to have qualified CSIR-UGC NET exam/DBT-JRF /DST INSPIRE fellowship etc. Project associate positions are advertised directly on the institute’s website.
Bachelor/Master/Thesis/Dissertation/Summer/Trainee:
Please contact the group leader directly by describing your motivation 2-3 months well before the desired start date.
International researchers:
If you ever had intentions to visit India, our laboratory is interested in hosting international researchers for both short term and long-term stays. Depending on the various funding opportunities applicable to individual applicants, the options can be mutually explored. Several programs such as DFG (German Research Foundation, Deutsche Forschungsgemeinschaft) postdoc fellowship, Erasmus traineeship, TWAS- DBT Ph.D. Scholarship for candidates from developing countries etc. allow hosting of international researchers in the group. Candidates are advised to contact the group leader directly to jointly figure out the possibilities.

